All cells make decisions in response to signals in their environment and their previous experiences. To divide or not to divide. To migrate towards or away from a substance. To conserve resources, or produce protective molecules. These decisions rest on the selective expression of gene products, a process orchestrated by gene regulation.
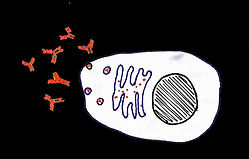

Cell division
Cell migration
Cell producing protective molecules
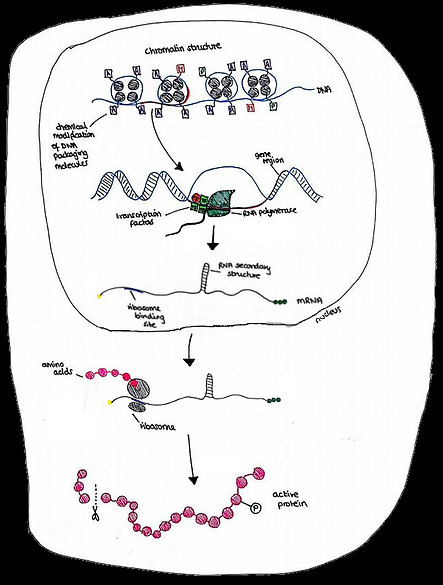
Gene regulation allows a cell to alter the species, timing and volume of proteins it produces, in response to environmental and intracellular signals. These proteins can then alter cell morphology and behaviour appropriately.
Gene regulation can occur at several steps in the process of gene expression. This includes the modification of DNA-packaging proteins, the binding of transcription factors, and the formation of secondary structures in mRNA.
The Central Dogma of Gene Expression in a eukaryotic cell:
DNA -> mRNA -> Protein
Transcription
Translation
Control of gene regulation is highly valued in the scientific community. Biomedical engineers, biologists, chemists and plant scientists working on gene therapy and genetically modified organisms need to control gene regulation to ensure that their modifications have appropriate effects.
So far, control has been achieved largely by hijacking cells’ intrinsic mechanisms, namely repurposing natural promoter sequences and their protein transcription factors.
For instance, insertion of a lac promoter before a gene allows the regulation of that gene in relation to the concentration of a chemical inducer called IPTG. IPTG causes unbinding of the repressor transcription factor molecule LacI, allowing gene transcription, and thereby gene expression, to occur.

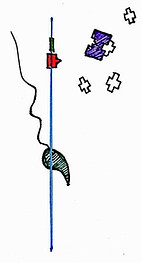


IPTG
LacI (transcription factor)

Our solution is elegant and effective: add multiple binding sites in conventional riboswitches for cooperative control of gene translation.
Well, let’s take it one step at a time. First we needed to ask if it were possible to make cooperative riboswitch structures.
Cooperativity requires that at least two chemical events are necessary to stably change from state one (e.g. OFF) to state two (e.g. ON). For positive cooperativity the first chemical event must be harder than the second.
In a cooperative switch, binding of the first trigger alters the switch molecule’s structure to make the second binding site more accessible or more complementary to the second trigger.
OUR SOLUTION (merit)
PROJECT GOALS (specification and feasibility)
Criteria for Success (Goals Checklist)
Core Goals
-
DESIGN
-
Design a novel cooperative activatory riboswitch
-
Model this riboswitch in silico: will it behave how we want it to?
-
-
IN VITRO TESTING
-
Characterise structure and folding: will it fold correctly in an in vitro system?
-
Characterise trigger strand binding behaviour: do the triggers bind to the switch and at what concentrations?
-
Model the riboswitch in silico with our data so far: what concentrations of riboswitch and trigger will we need to test switching behaviour?
-
Characterise switching behaviour in the in vitro system: is it cooperative?
-
Characterise the switching behaviour of a riboswitch with two different binding domain sequences: is trigger binding sequence-specific?
-
Using our in vitro data, model the behaviour of the switch with two different binding domain sequences in silico: can we calculate the Kd of trigger binding?
-
Characterise switching behaviour of riboswitches with shorter and longer stem lengths: is it tunable?
-
-
IN SILICO TESTING
-
Develop a framework for measuring cooperativity:
-
Can we use hill curves?
-
Can we compare different types of cooperative system?
-
-
Create a mechanistic model of each main switch design
- Use the models to fit to and explain experimental data
- Demonstrate fine tuning of the switching threshold and level of cooperativity
-
-
IN VIVO TESTING
-
Transform the bacteria: can we get our plasmids into the cells?
-
Measure the red fluorescence of...
-
untransformed cells: can we confirm that cells without our plasmids don't produce red fluorescent protein (negative control)?
-
transformed cells that don't contain our activatory riboswitch: is the fluorescent protein expressed (positive control)?
-
transformed cells that do contain our activatory riboswitch: will the riboswitch (in it's OFF state) suppress translation in vivo?
-
transformed cells that contain activatory riboswitches with longer or shorter stems in their OFF states: do longer stems suppress translation more (is our riboswitch tunable in vivo)?
-
transformed cells (that contain our activatory riboswitch) with increasing concentrations of arabinose: will the riboswitch turn ON cooperatively in vivo in response to increasing concentrations of trigger expression?
-
-
Troubleshooting: measure the green fluorescence of...
-
untransformed cells: can we confirm that cells without our plasmids don't produce green fluorescing triggers (negative control)?
-
transformed cells that contain our triggers, with increasing concentrations of arabinose: are the triggers expressed and folding correctly? ...ah, here's our problem.
-
-
Stretch Goals
-
Use single cell analysis and flow cytometry to characterise switch behaviour in vivo: can we investigate switch behaviour in vivo on a single cell scale?
- Design other novel cooperative activatory riboswitches
-
Homotropic asymmetric design
-
Double toehold design
-
Triple trigger design
-
-
Design a novel cooperative repressive riboswitch
-
Characterise repressive riboswitch folding in vitro: does the switch remain in the ON state without triggers?
-
Characterise trigger strand binding behaviour in vitro: will the switch bind the triggers as designed?
-
Characterise repressive riboswitch switching behaviour in vitro: does the switch turn OFF cooperatively?
-
Transform bacteria: can we get our plasmids into cells?
-
Measure red fluorescence: is the fluorescent protein expressed when our switch is in the ON state (no triggers)?
-
Add arabinose to increase trigger expression and measure red and green fluorescence: are our triggers expressed and folding correctly, and do they turn the switch OFF in a cooperative manner (reduce red fluorescence)?
-
The ideal outcome of this project
The ideal outcome for this project is to design, synthesise, and verify a novel activatory riboswitch which can regulate gene expression in a cooperative, tunable, predictable and modular manner. If we can make additional activatory riboswitches, or even repressive riboswitches, so much the better!
In succeeding, we will have developed a new molecular gene regulatory tool to be used in the construction of gene circuits for biocomputers.
Our goals for BIOMOD
Given the time constraints of the BIOMOD project, we decided to break our ideal outcome down into a set of smaller sequential milestones that we could build on in future. We chose three core ‘proof of concept’ goals that would support the future development of this technology.
-
Demonstrate that our activatory designs fold correctly and function cooperatively in vitro
-
Demonstrate that our activatory designs are tunable in vitro
-
Demonstrate cooperative switch function in vivo
To achieve these points, we had to make alterations to our designs to make them more testable:
-
Our in vitro switch designs were tested with ssDNA rather than RNA. ssDNA has similar properties to RNA but is much cheaper so we could order more strands and test a greater variety of designs.
-
The in vitro switches were tested in isolation using fluorophore-quencher pairs rather than within the full context of the gene
-
We tested our switches in vivo in a laboratory E. coli strain using fluorescent reporter genes to measure gene translation rate, and fluorescent 'broccoli' aptamer constructs to measure trigger concentration. I.e. we used a simplified and well-understood model system, but one that unfortunately won't be applicable to every application of our switch.
FUTURE WORK
The purpose of this project isn’t to develop gene regulatory biocomputing technology with any direct diagnostic or medicinal value. We want to prove that cooperative riboswitches can be made. In future, they could potentially be modified by changing the complementary site sequences to detect pathological RNA sequences, and placed so that they regulate the expression of medicinal genes, but this is outside the scope for our current project.
We envisage several interesting avenues for future work on our cooperative riboswitches, including...
-
Increasing the number of cooperative sites on the switches to enhance their cooperativity and switch-like behaviour further.
-
Implementing a "ribo-attenuator" element to help insulate the switches from contextual changes, allowing the switches to regulate any protein and become truly modular.
-
Investigating other variables affecting switch activation and tunability e.g. trigger length, ribosome binding site strength and ribosome binding site position.
-
Incorporating multiple switches into one system to demonstrate independent regulation of several genes.
-
Changing the Broccoli aptamer to other RNA aptamers, e.g. an aptamer that binds to a downstream product to create a feedback loop. This would mean that the gene's protein output would regulate its rate of mRNA translation, thus altering the amount of protein produced - eventually a steady state of protein quantity could be reached.
A really good riboswitch for use in gene therapy or logic-gated biocomputing would have four qualities. It would be:
-
Predictable
-
Modular, with few sequence limitations
-
Cooperative
-
Tunable
As shown above, effective riboswitches have already been designed, but so far none have any cooperative behaviour. This means that although they can regulate gene expression, they don’t display sharp switch-like on-off behaviour that would be most useful in biocomputing and other applications.
What if we could design a modular cooperative activatory gate riboswitch? So that when we add the trigger, it displays the switch-like on-off behaviour that we want. Could we alter the sensitivity of the switch and range of the response? Could we make it tunable? And will it work in cells where everything gets more compressed and complicated?
THE PROBLEM (relevance)
Project
BACKGROUND (relevance)
All cells make decisions in response to signals in their environment and their previous experiences. To divide or not to divide. To migrate towards or away from a substance. To conserve resources, or produce protective molecules. These decisions rest on the selective expression of gene products, a process orchestrated by gene regulation.


Cell division
Cell migration
Cell producing protective molecules
Gene regulation allows a cell to alter the species, timing and volume of proteins it produces, in response to environmental and intracellular signals. These proteins can then alter cell morphology and behaviour appropriately.
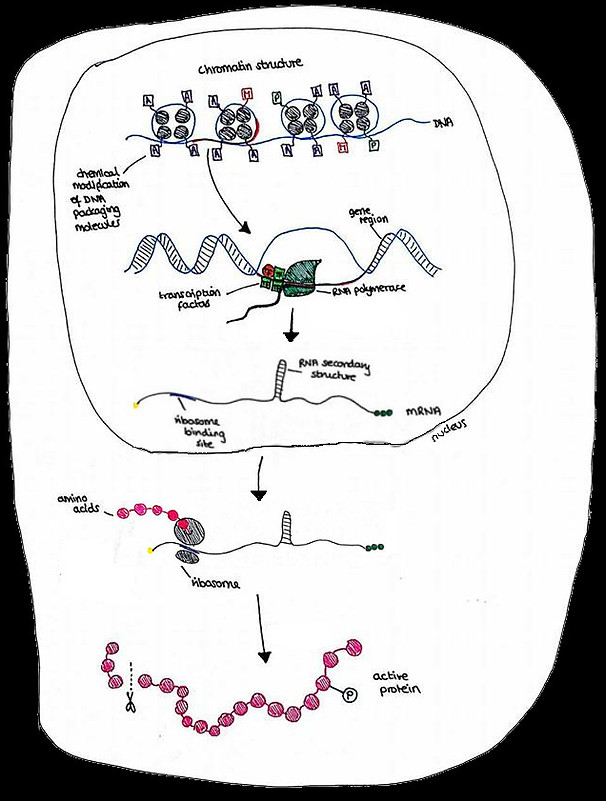
Gene regulation can occur at several steps in the process of gene expression. This includes the modification of DNA-packaging proteins, the binding of transcription factors, and the formation of secondary structures in mRNA.
The Central Dogma of Gene Expression in a eukaryotic cell:
DNA -> mRNA -> Protein
Transcription
Translation
Control of gene regulation is highly valued in the scientific community. Biomedical engineers, biologists, chemists and plant scientists working on gene therapy and genetically modified organisms need to control gene regulation to ensure that their modifications have appropriate effects.
So far, control has been achieved largely by hijacking cells’ intrinsic mechanisms, namely repurposing natural promoter sequences and their protein transcription factors.
For instance, insertion of a lac promoter before a gene allows the regulation of that gene in relation to the concentration of a chemical inducer called IPTG. IPTG causes unbinding of the repressor transcription factor molecule LacI, allowing gene transcription, and thereby gene expression, to occur.




IPTG (chemical inducer)
LacI (transcription factor)
Operator
RNA polymerase
Lac promoter
Bound by LacI, expression is repressed
IPTG binds LacI, releasing the operator
RNA synthesis occurs -> gene expression
Illustration of Lac promoter gene regulation
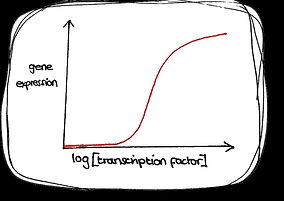
Cooperative
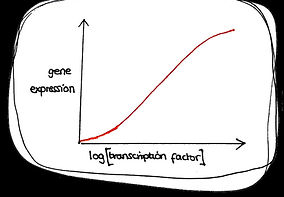
Non cooperative
DNA
mRNA
Other promoters can couple gene expression to the growth conditions of the cell, its position in the cell cycle, or to chemical modifications of DNA-packaging proteins. Protein-protein interactions between transcription factors can also display cooperativity, creating useful concentration-dependent switch-like behaviours in cells.
However, simple hijacking at the level of mRNA transcription has many limitations.
Cells are perfectly designed to function in their natural environments. Many of their gene regulation pathways interact, affecting the activation and repression of diverse groups of related genes at the same time. The inducers are molecules found in the cell’s environment, like lactose, and the regulation pathways and decisions can change depending on the presence of many different inputs.
This network of interactions is optimal for the cell, but not so useful for the scientist trying to reliably control just one gene in the network, let alone the synthetic biologist trying to build a biocomputer with many independently-regulated genes.

As the domain sequences used are not greatly constricted in the domain-based riboswitches above, scientists could use several of these riboswitches simultaneously to regulate several genes independently of one another, isolating the effects of one gene or even building logic-gated biocomputers.
Logic-gated biocomputers are genetically-modified cells that can detect and respond as we dictated to external stimuli. They might one day allow us to detect disease and deliver medicine from within the body itself. For example, by creating a regulatory switch pathway to induce insulin gene expression in response to glucose-concentration, a cell-based treatment for diabetes could be developed. Logic-gating of toxin gene expression to express cell-destroying drugs in malignant cells only could allow the development of a new gene therapy for cancer.
Network of genes
One solution is to make new, synthetic protein transcription factor systems. Unfortunately, they are incredibly difficult to design. Protein folding, and thereby its interactions with DNA and other proteins, cannot be reliably predicted from its amino acid sequence, even with the supercomputing technology currently available. Therefore, designing new transcription factors and promoters would take a lot of resources, a long time, and a lot of trial and error. At the moment, it’s just not feasible.
RNA, on the other hand, folds in a more predictable manner.
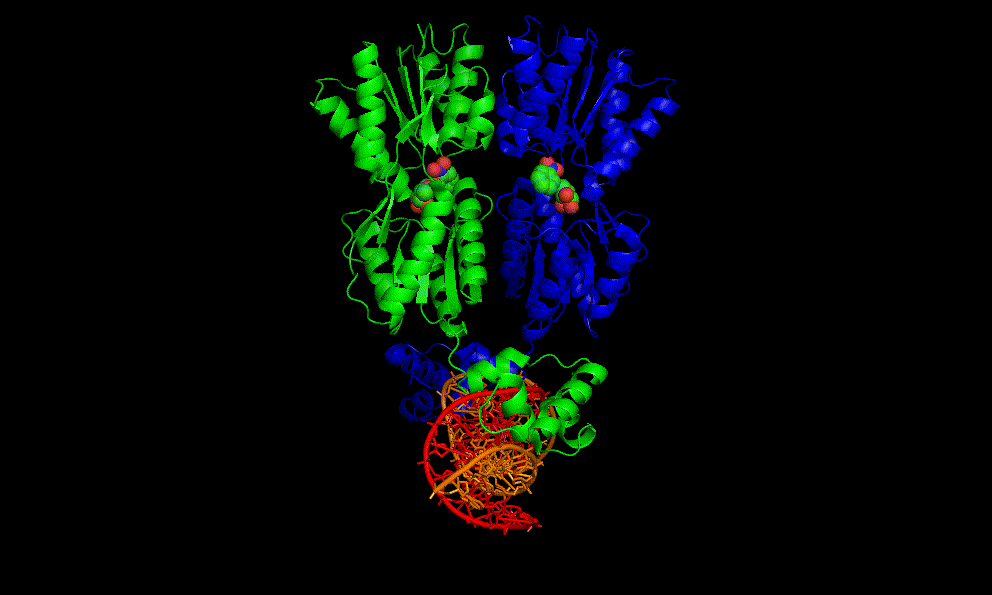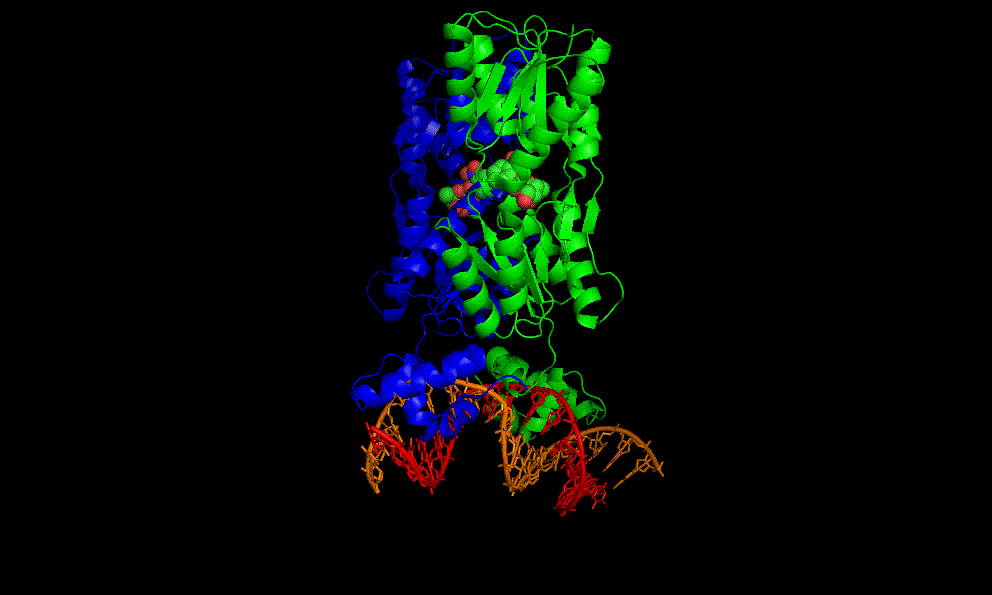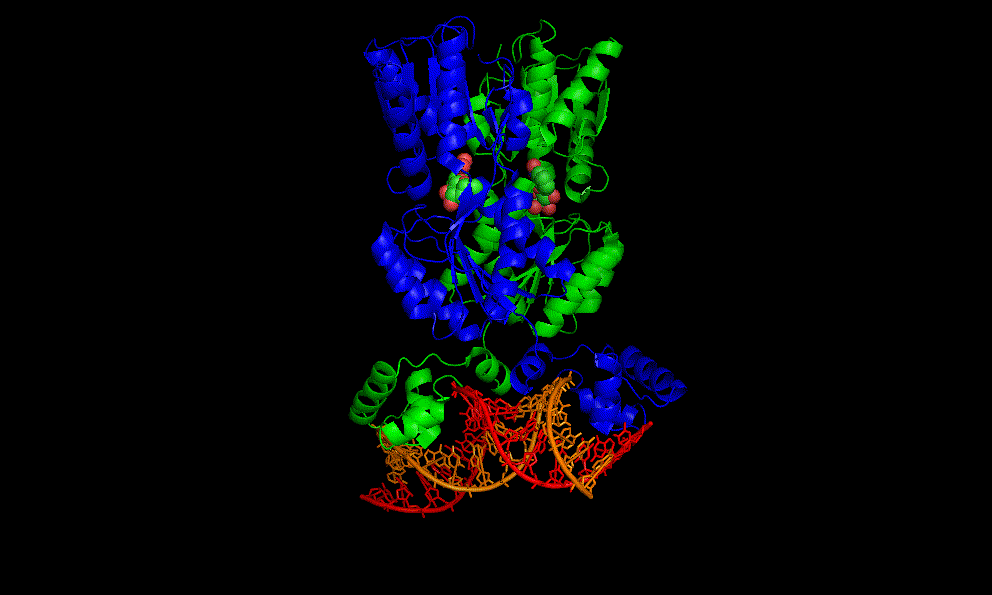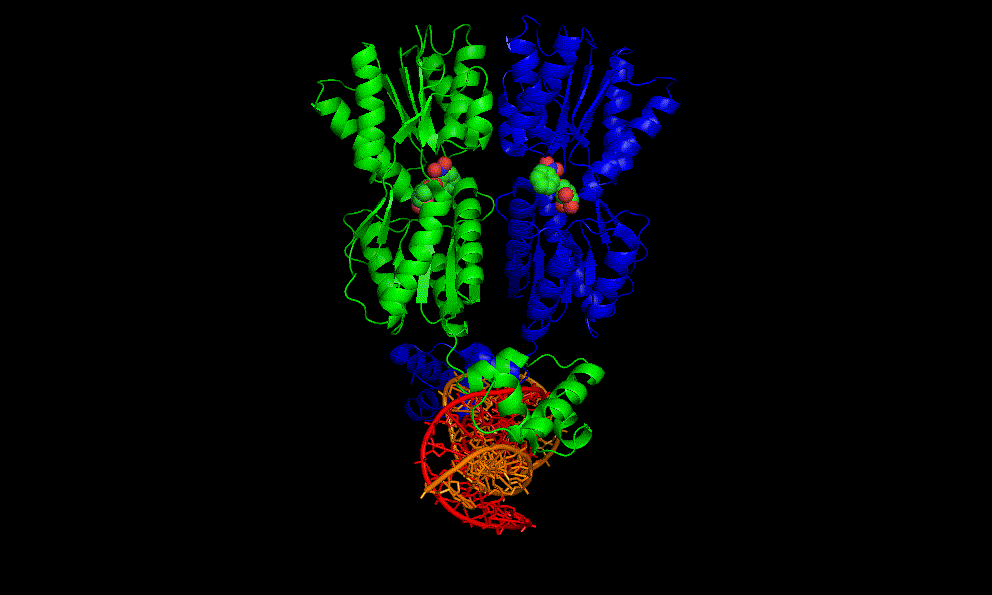
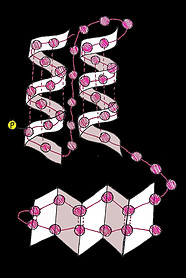
The flavin mononucleotide (FMN) riboswitch


20 different amino acids
[1' structure]
4 different nucleotide bases
[1' structure]
Chemical modification of amino acids
Protein transcription factor on DNA
=> complex folding behaviour
RNA hairpin occluding ribosome binding site
=> more predictable folding behaviour
RNA can fold via complementary base-pairing rules to form loops, twists and hairpins from a previously disordered chain. If these structures happen to restrict the ribosomes access to a ribosome binding site then the ribosome will be prevented from binding to the mRNA and gene expression will be reduced.
Riboswitches are RNA structures containing aptamers: sequences of bases that are able to bind to other molecules such as amino acids or metal ions. When these molecules bind to the aptamer sequence it causes a structural change in the riboswitch that affects gene expression.
Bacteria, like E. coli, commonly use riboswitches to control genes involved in their metabolism. One of these is the FMN riboswitch illustrated below.
hydrogen bonding (almost) always A-T, C-G (complementary bases)
[2' structure]
hydrogen bonding between any combination of amino acids forms a-helices and b-pleated sheets
[2' structure]
Interactions between 2' structures [3' structure] and between amino acid chains [4' structure]
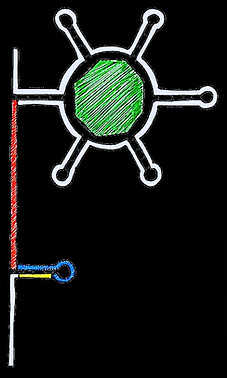
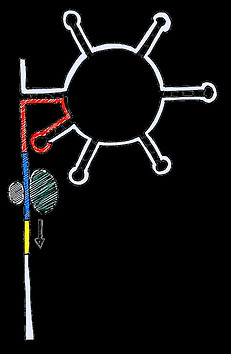
ON
AUG
OFF
RBS occluded by hairpin loop
FMN
aptamer
AUG
RBS can be recognised by ribosome
5'
5'
3'
3'
gene
gene
Two more examples of natural riboswitches are illustrated below.
Some riboswitches are activatory, i.e. gene expression increases when then the molecule is added. Others are repressive where the molecule reduces expression. Some are reversible, allowing quick responses to both the addition and removal of a trigger, whereas others, like the self-cleavage riboswitch, cause degradation of the mRNA preventing gene expression until more mRNA is synthesised.
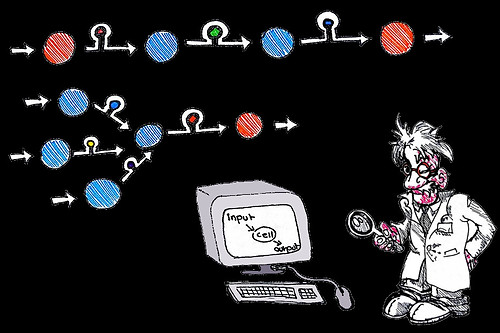
Glucose detection
Insulin production
Tumour markers
Toxin production
Bioengineered riboswitches between genes
Simplified logic-gated biocomputer
What if we could take it a step further? Though harder to design, a cooperative repressive riboswitch would have many more applications than an activatory riboswitch in biocomputing. Could we make a cooperative, modular riboswitch that stops translation when triggers are added?
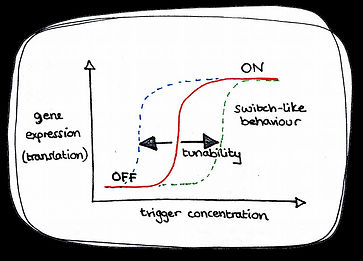
Behaviour of a tunable cooperative activatory riboswitch

Could this be the breakthrough in logic-gated biocomputing? And one day, could riboswitches help programme synthetic cells to autonomously detect and treat disease, without ever needing the input of a medical team?
Behaviour of a tunable cooperative repressive riboswitch
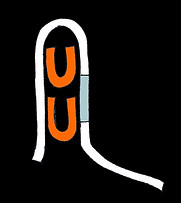
Our repressive switch is disordered when there are no triggers around. With it's exposed ribosome binding site, translation can to occur. When triggers are present in high enough concentrations, they will preferentially bind at the two complementary sites nearest one another, pulling the long switch strand into a paperclip-like shape. This places the second set of complementary sites closer together, making it easier for the second trigger to bind. The second trigger occludes the ribosome binding site, shutting off translation and stabilising the structure.
Our activatory
switch mechanism
Our activatory switch is a stem-loop structure with the ribosome binding site occluded in the stem.
Two short RNA triggers can bind to the loop, overlapping with a part of the stem. The binding of the first trigger is difficult because binding requires the stem to break and the loop to straighten. However breaking the stem makes it easier for the second trigger to also bind.
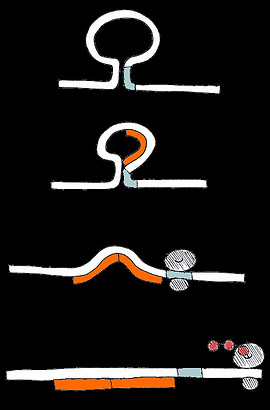
One trigger strand binds, straining and breaking the stem
A second trigger strand binds, stabilising the open, ON state with its exposed ribosome binding site
The ribosome can recognise the ribosome binding site and begin translation
The closed, OFF state

Activatory switch
5'
3'
ribosome binding site
gene
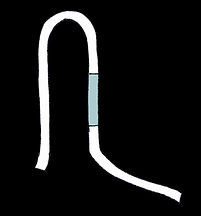
Repressive switch
5'
3'
ribosome binding site
gene
These designs have many advantages. Our riboswitches represent modular, tunable synthetic molecular tools to predictably and cooperatively regulate gene expression.
They are simple and small, and placed before the gene sequence, reducing the likelihood that they will fold incorrectly. Our riboswitches are also designed to be modular – they don’t need to interact with any part of the gene sequence after the ribosome binding site, so they can theoretically be used to regulate any transcribed gene.
Because they have few sequence restrictions, many different versions of the same designs could be used within the same cell, regulating genes independently of one another.
Simple characteristics, like stem length and loop length, can also be altered to change their sensitivity, tuning them.
Our repressive switch mechanism


The open, ON state
One trigger strand binds, clipping the switch into a loop
A second trigger binds, stabilising the clipped conformation and occluding the ribosome binding site, halting translation
After many days in the computer room reading widely, sketching poorly, and plugging code into Nupack with varying success, we came up with some pretty elegant designs (though we do say so ourselves).
Our solutions are innovative because they combine synthetic riboswitches and cooperativity: two existing ideas that have never been explored together. The designs are elegant as they only require simple changes to conventional riboswitches to generate the useful and interesting effects of cooperativity.

A cooperative reaction
Lastly, their clear cooperative mechanisms allow them to display the switch-like behaviour necessary to function as logic gates.
Creation of biocomputers aside, the development of synthetic gene regulation tools has proven to be a challenge in biosynthetic technology. This project offers a novel solution. It also offers opportunities for the ongoing research of DNA folding behaviour, including the testing of current in silico models.
Further details about our activatory and repressive switches as well as the process we used to come up with our designs can be found in our design pages. To learn how our designs operated in our testing please see our results pages.
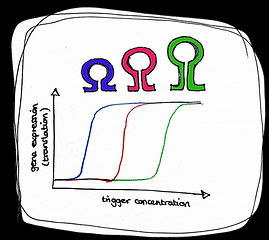
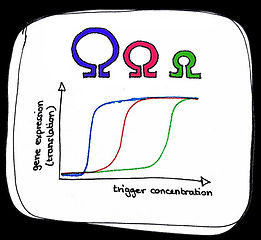
Stem length alters sensitivity
Loop length alters sensitivity
Within the time limits of the BIOMOD competition, it’s important to choose goals that are both achievable and scientifically valuable.
The design of novel modular, tunable and cooperative riboswitches could be taken in many different interesting and rewarding directions. For this project, we felt that it was important to choose strategic goals that allowed us to test the core concept of cooperativity within our designs, and their tunability. We also decided to focus largely on our activatory riboswitch, as we felt that it had the greatest likelihood of success.
There are two major aspects to our project that made us confident that our project would be feasible:
1) Past Precedent
Non-cooperative de-novo-designed synthetic riboswitches have not only been constructed, but successfully incorporated and tested in genetic circuits. Green et al. (2014) showed that it was possible to incorporate their forward-engineered toehold riboswitches into prokaryotic cells, exploiting the specificity of RNA sequences to control 12 genes independently. Simon et al. (2014) demonstrated that the rational design of cooperative molecular constructs (not riboswitches) was possible. Together, their work suggested that our project was not only scientifically and technically interesting, but also feasible.
2) Project Design
We designed our project to generate initial results quickly. For example, we tested the architecture of our riboswitches in vitro using quick assays like mixing strands and measuring fluorescence quenching. Once we knew which designs were most likely to work, we could prioritise these in the more time-consuming steps of cell transformation and in vivo testing.


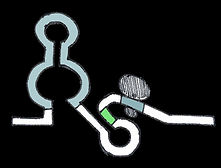

Small molecule-triggered repressive riboswitch
Small molecule binding refolds the mRNA, allowing occlusion of the ribosome binding site and thereby halting translation

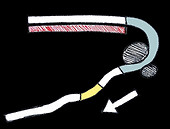
RNA toehold-triggered activatory riboswitch
A complementary single stranded RNA trigger binds the partially exposed part of the domain. This then breaks the occlusive stem bonds of the stem-loop structure, exposing the ribosome binding site, and thereby allowing translation to occur.

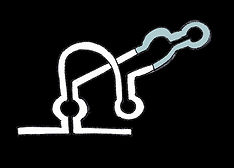
Ligand-triggered self-cleavage riboswitch
Ligand binding draws communicating regions into close proximity, allowing autocatalytic cleavage and degradation of the mRNA, halting translation
Scientists have experimented with natural riboswitches discovered in many bacterial species and have also developed new synthetic riboswitches.
These riboswitches are domain-based like the "toehold" switch illustrated below. These use RNA sequences, called domains, that can bind to complementary RNA sequences both within the riboswitch structure and on separate RNA molecules called trigger RNAs.
binding event 1
binding event 2
E1 > E2
|G1| < |G2|
These compromises allowed us to test the design sequences in a more efficient manner. Once successful, we could move on to using more expensive materials.

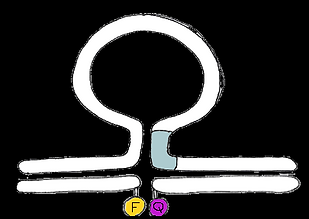
Testing our switch in vitro
fluorophore-quencher pair attached to hybridising ssDNA strands
ssDNA rather than RNA
In the OFF state, the quencher molecule is in close proximity to the fluorophore, reducing fluorescence emission.
In the ON state, the quencher molecule and fluorophore are far apart, so fluorescence emission is high.
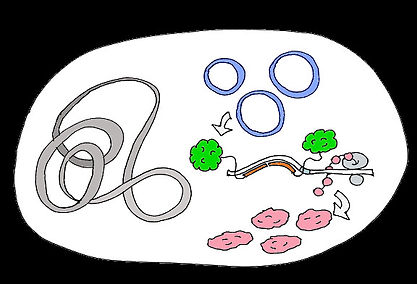
Testing our switch in vivo
plasmids containing our constructs
fluorescent green 'broccoli' aptamer attached to triggers
red fluorescent protein produced on gene translation (when the switch is in the ON state)
native E. coli DNA
E. coli cell